
Main Functions
main_functions.RmdIntroduction
Outline
In this vignette we explain how to use the package’s main functions. Main functions are functions that are part of the minimal workflow of data preparation for IRT analyses using item meta data, that is, performing checks, merging, recoding, aggregating and scoring variables. For explanation of further important functions and diagnostic tools for data preparation and plausibility checks, there will be another vignette.
You can click on the specific function to jump to its explanation.
| Function | Explanation |
|---|---|
| Reading in (Meta) Data | |
| readDaemonXlsx() | read in the inputlist that was created using the EDV-tool ‘ZKDaemon’. |
| readSpss() | read in SPSS files. |
| readMerkmalXlsx() | read in additional item and exercise attributes like processing time, formats, content categories, … |
| Checks | |
| checkInputList() | check the inputList for internal consistency. |
| checkData() | check data sets according to item meta information and other plausibility checks of the data. |
| checkDesign() | check data sets according to test design meta information. |
| Merging, Recoding, Aggregating, Scoring | |
| mergeData() | merging the data sets and diagnostics to ensure a fit. |
| recodeData() | recode the subitems according to meta information from the inputList. |
| aggregateData() | aggregate subitems into items. |
| scoreData() | recode items that previously consisted out of subitems. |
| mnrCoding() | recoding the last items (if empty) in each block (see test design) as ‘missing not reached’. |
| Wrapper | |
| automateDataPreparation() | wraps most of the other features into one big function. |
| Additional Diagnostics and Rater Tools | |
| catPbc() | calculates category-level point-biserial correlations for recoded and raw item data. |
| evalPbc() | flags suspicious category discrimination patterns from
the output of catPbc(). |
| meanAgree() | calculates mean pairwise percentage agreement across raters. |
| meanKappa() | calculates mean pairwise kappa across raters. |
| make.pseudo() | reduces multiple real raters to a smaller number of pseudo raters. |
| computeCutsIDM() | computes cut scores for Item Descriptor Matching based on monotonized moving averages. |
| plotCutsIDM() | plots the raw ratings, moving averages, monotonized
curves and cut scores from computeCutsIDM(). |
| visualSubsetRecode() | supports interactive visual inspection and recoding of flagged subsets. |
| Export | |
| collapseMissings() | recodes missing types into predefined scores (usually 0, 1, and NA). Such a collapsed R data frame can be passed directly to eatModel for scaling. |
| writeSpss() | produces an SPSS syntax and a .txt data set that can be read into SPSS with the syntax including all meta data. |
| prep2GADS() | both the raw data sets and the finished, scored data sets, including all their meta data, can be exported into a GADSdat object for data storage or further processing in eatGADS. |
Data Structure
In the Getting Started Vignette we
saw what we use the eatPrep package for and how the data
structure looks like. Now, we will look at the main functions to learn
how to implement that via R.
As a reminder, an eatPrep data set contains the
following layers:
| booklets | containing blocks |
| blocks | containing units (= items) |
| units | containing subunits/subitems |
| subunits | having values and |
| values | including missings and recode values |
And here is an overview for different value types the IQB uses:
| Code | Label | Abbr | Explanation |
|---|---|---|---|
| -98 | mir | missing invalid response | (1) Item was edited, and (2a) empty answer or (2b) invalid (joke) answer. |
| -99 | mbo | missing by omission | Item wasn’t edited but seen, or wasn’t seen, but there are seen or edited subsequent Items. |
| -96 | mnr | missing not reached | (1) Item wasn’t seen, and (2) all subsequent Items weren’t seen, either. |
| -97 | mci | missing coding impossible | (1) Item should/could have been edited, and (2) answer can’t be analysed due to technical problems. |
| -94 | mbd | missing by design | no answer, because Item wasn’t shown to the testperson by design. |
Input
inputMinimal contains the bare minimum for all functions
in eatPrep to work, inputList contains
additional meta data about the items (such as information about the
task’s content).
For more information on how the different layers or input tables look, please see the Getting Started Vignette.
inputMinimal <- list(units = inputList$units[ -nrow(inputList$units), c("unit", "unitAggregateRule")],
subunits = inputList$subunits[, c("unit", "subunit","subunitRecoded")],
values = inputList$values[ , c("subunit", "value", "valueRecode", "valueType")],
unitRecodings = inputList$unitRecodings[ , c("unit", "value", "valueRecode", "valueType")],
blocks = inputList$blocks,
booklets = inputList$booklets,
rotation = inputList$rotation)Input Data
We have several overlapping booklets with several blocks in each booklet. Moreover, there is a unique identifier for each person and some additional information about each student like their gender or their socioeconomic status. There is one data set per booklet. In order to prepare the data, we need to construct one large data set.
In order to do that we first need to read the data into R, check the
data for invalid or incorrect codes and then merge the data into one
data set. In the following the different functions to do that are
described. inputDat gives us a first idea on how the data
is supposed to look like.
# looking at the data
str(inputDat)
#> List of 3
#> $ booklet1:'data.frame': 100 obs. of 25 variables:
#> ..$ ID : chr [1:100] "person100" "person101" "person102" "person103" ...
#> ..$ hisei: num [1:100] 49 NA 57 32 59 56 55 47 NA 50 ...
#> ..$ I01 : chr [1:100] "1" "9" "3" "2" ...
#> ..$ I02 : chr [1:100] "4" "4" "4" "4" ...
#> ..$ I03 : chr [1:100] "1" "2" "2" "3" ...
#> ..$ I04 : chr [1:100] "2" "2" "3" "1" ...
#> ..$ I05 : chr [1:100] "0" "0" "0" "1" ...
#> ..$ I06 : chr [1:100] "0" "9" "0" "1" ...
#> ..$ I07 : chr [1:100] "2" "2" "2" "3" ...
#> ..$ I08 : chr [1:100] "0" "1" "0" "0" ...
#> ..$ I09 : chr [1:100] "2" "2" "2" "3" ...
#> ..$ I10 : chr [1:100] "2" "1" "1" "4" ...
#> ..$ I11 : chr [1:100] "4" "1" "3" "2" ...
#> ..$ I12a : chr [1:100] "1" "0" "1" "0" ...
#> ..$ I12b : chr [1:100] "0" "9" "0" "1" ...
#> ..$ I12c : chr [1:100] "4" "1" "2" "4" ...
#> ..$ I13 : chr [1:100] "9" "4" "9" "4" ...
#> ..$ I14 : chr [1:100] "9" "4" "9" "1" ...
#> ..$ I15 : chr [1:100] "9" "3" "9" "1" ...
#> ..$ I16 : chr [1:100] "9" "2" "9" "2" ...
#> ..$ I17 : chr [1:100] "9" "3" "9" "4" ...
#> ..$ I18 : chr [1:100] "9" "1" "9" "1" ...
#> ..$ I19 : chr [1:100] "9" "4" "9" "2" ...
#> ..$ I20 : chr [1:100] "9" "1" "9" "1" ...
#> ..$ I21 : chr [1:100] "9" "2" "9" "3" ...
#> $ booklet2:'data.frame': 100 obs. of 25 variables:
#> ..$ ID : chr [1:100] "person200" "person201" "person202" "person203" ...
#> ..$ hisei: num [1:100] 69 76 47 58 62 78 70 26 57 70 ...
#> ..$ I08 : chr [1:100] "0" "0" "1" "0" ...
#> ..$ I09 : chr [1:100] "2" "4" "1" "2" ...
#> ..$ I10 : chr [1:100] "3" "1" "2" "2" ...
#> ..$ I11 : chr [1:100] "4" "2" "1" "4" ...
#> ..$ I12a : chr [1:100] "1" "0" "1" "1" ...
#> ..$ I12b : chr [1:100] "1" "0" "1" "1" ...
#> ..$ I12c : chr [1:100] "4" "3" "2" "2" ...
#> ..$ I13 : chr [1:100] "4" "2" "1" "2" ...
#> ..$ I14 : chr [1:100] "3" "4" "2" "3" ...
#> ..$ I15 : chr [1:100] "4" "1" "4" "4" ...
#> ..$ I16 : chr [1:100] "4" "3" "3" "2" ...
#> ..$ I17 : chr [1:100] "4" "1" "2" "3" ...
#> ..$ I18 : chr [1:100] "1" "4" "3" "2" ...
#> ..$ I19 : chr [1:100] "2" "1" "4" "3" ...
#> ..$ I20 : chr [1:100] "4" "9" "2" "2" ...
#> ..$ I21 : chr [1:100] "2" "9" "2" "1" ...
#> ..$ I22 : chr [1:100] "1" "9" "2" "1" ...
#> ..$ I23 : chr [1:100] "9" "9" "2" "3" ...
#> ..$ I24 : chr [1:100] "1" "9" "3" "3" ...
#> ..$ I25 : chr [1:100] "1" "9" "0" "0" ...
#> ..$ I26 : chr [1:100] "9" "9" "1" "0" ...
#> ..$ I27 : chr [1:100] "0" "9" "1" "0" ...
#> ..$ I28 : chr [1:100] "1" "9" "1" "0" ...
#> $ booklet3:'data.frame': 100 obs. of 23 variables:
#> ..$ ID : chr [1:100] "person300" "person301" "person302" "person303" ...
#> ..$ hisei: num [1:100] 49 NA 57 32 59 56 55 47 NA 50 ...
#> ..$ I01 : chr [1:100] "2" "3" "2" "1" ...
#> ..$ I02 : chr [1:100] "4" "3" "3" "3" ...
#> ..$ I03 : chr [1:100] "3" "1" "1" "1" ...
#> ..$ I04 : chr [1:100] "1" "3" "1" "1" ...
#> ..$ I05 : chr [1:100] "0" "1" "1" "0" ...
#> ..$ I06 : chr [1:100] "9" "1" "0" "1" ...
#> ..$ I07 : chr [1:100] "8" "2" "2" "1" ...
#> ..$ I15 : chr [1:100] "3" "9" "9" "9" ...
#> ..$ I16 : chr [1:100] "1" "3" "3" "3" ...
#> ..$ I17 : chr [1:100] "4" "4" "4" "4" ...
#> ..$ I18 : chr [1:100] "4" "4" "4" "4" ...
#> ..$ I19 : chr [1:100] "2" "9" "9" "9" ...
#> ..$ I20 : chr [1:100] "3" "4" "2" "4" ...
#> ..$ I21 : chr [1:100] "3" "2" "2" "2" ...
#> ..$ I22 : chr [1:100] "3" "4" "4" "9" ...
#> ..$ I23 : chr [1:100] "3" "1" "1" "9" ...
#> ..$ I24 : chr [1:100] "1" "1" "1" "9" ...
#> ..$ I25 : chr [1:100] "0" "1" "9" "9" ...
#> ..$ I26 : chr [1:100] "1" "0" "9" "9" ...
#> ..$ I27 : chr [1:100] "0" "9" "9" "9" ...
#> ..$ I28 : chr [1:100] "0" "9" "9" "9" ...Reading in (Meta) Data
First, we need to get the information from our IQB data bank via ZKDaemon, or alternatively via SPSS.
Items via ZKDaemon
ZKDaemon is a program used by IQB that can be found in the IQB internal folders (i:). After installing you can get the meta data and the items from your specific study using information stored in the IQB Databases (DB2/DB3/DB4). Alternatively you can get the meta data from an SPSS file via ZKDaemon. Within the program you can set missing types and import a test design. Then you can produce Excel files, which are expected to have the following sheets: “units”, “subunits”, “values”, “unitrecoding”, “sav-files”, “params”, “aggregate-missings”, “itemproperties”, “propertylabels”, “booklets”, and “blocks”.
The function readDaemonXlsx() reads the Excel file into
R and will produce a warning if any sheets are missing. It needs one
character string (filename) containing path, name and extension of the
Excel file (.xlsx) produced by ZKDaemon.
The function returns a list of data frames containing information
that is required by the data preparation functions.
inputList shows an example of this list.
filename <- system.file("extdata", "inputList.xlsx", package = "eatPrep")
readDaemon <- readDaemonXlsx(filename)
#> Reading sheet 'units'.
#> Reading sheet 'subunits'.
#> Reading sheet 'values'.
#> Reading sheet 'unitrecoding'.
#> Reading sheet 'sav-files'.
#> Reading sheet 'params'.
#> Reading sheet 'aggregate-missings'.
#> Reading sheet 'booklets'.
#> Reading sheet 'blocks'.
str(readDaemon)
#> List of 9
#> $ units :'data.frame': 29 obs. of 6 variables:
#> ..$ unit : chr [1:29] "I01" "I02" "I03" "I04" ...
#> ..$ unitLabel : chr [1:29] "Animals: Weight of a duck" "Animals: Weight of a horse" "Animals: Weight of a mouse" "Animals: Weight of a cat" ...
#> ..$ unitDescription : chr [1:29] NA NA NA NA ...
#> ..$ unitType : chr [1:29] "TI" "TI" "TI" "TI" ...
#> ..$ unitAggregateRule: chr [1:29] NA NA NA NA ...
#> ..$ unitScoreRule : chr [1:29] NA NA NA NA ...
#> $ subunits :'data.frame': 30 obs. of 9 variables:
#> ..$ unit : chr [1:30] "I01" "I02" "I03" "I04" ...
#> ..$ subunit : chr [1:30] "I01" "I02" "I03" "I04" ...
#> ..$ subunitType : chr [1:30] "1" "1" "1" "1" ...
#> ..$ subunitLabel : chr [1:30] "Animals: Weight of a duck" "Animals: Weight of a horse" "Animals: Weight of a mouse" "Animals: Weight of a cat" ...
#> ..$ subunitDescription : chr [1:30] NA NA NA NA ...
#> ..$ subunitPosition : chr [1:30] "a)" "b)" "c)" "d)" ...
#> ..$ subunitTransniveau : chr [1:30] NA NA NA NA ...
#> ..$ subunitRecoded : chr [1:30] "I01R" "I02R" "I03R" "I04R" ...
#> ..$ subunitLabelRecoded: chr [1:30] "Recoded Animals: Weight of a duck" "Recoded Animals: Weight of a horse" "Recoded Animals: Weight of a mouse" "Recoded Animals: Weight of a cat" ...
#> $ values :'data.frame': 220 obs. of 8 variables:
#> ..$ subunit : chr [1:220] "I01" "I01" "I01" "I01" ...
#> ..$ value : chr [1:220] "1" "2" "3" "6" ...
#> ..$ valueRecode : chr [1:220] "0" "0" "1" "mnr" ...
#> ..$ valueType : chr [1:220] "vc" "vc" "vc" "mnr" ...
#> ..$ valueLabel : chr [1:220] "Response option 1 marked" "Response option 2 marked" "Response option 3 marked" "missing not reached" ...
#> ..$ valueDescription : chr [1:220] "Response option 1 marked" "Response option 2 marked" "Response option 3 marked" "missing not reached" ...
#> ..$ valueLabelRecoded : chr [1:220] "0" "0" "1" "mnr" ...
#> ..$ valueDescriptionRecoded: chr [1:220] NA NA NA NA ...
#> $ unitRecodings:'data.frame': 7 obs. of 7 variables:
#> ..$ unit : chr [1:7] "I12" "I12" "I12" "I12" ...
#> ..$ value : chr [1:7] "0" "1" "2" "3" ...
#> ..$ valueRecode : chr [1:7] "0" "0" "0" "1" ...
#> ..$ valueType : chr [1:7] "vc" "vc" "vc" "vc" ...
#> ..$ valueLabel : chr [1:7] NA NA NA NA ...
#> ..$ valueDescription : chr [1:7] NA NA NA NA ...
#> ..$ valueLabelRecoded: chr [1:7] NA NA NA NA ...
#> $ savFiles :'data.frame': 3 obs. of 3 variables:
#> ..$ filename: chr [1:3] "booklet1.sav" "booklet2.sav" "booklet3.sav"
#> ..$ case.id : chr [1:3] "ID" "ID" "ID"
#> ..$ fullname: chr [1:3] NA NA NA
#> $ newID :'data.frame': 1 obs. of 2 variables:
#> ..$ key : chr "master-id"
#> ..$ value: chr "ID"
#> $ aggrMiss :'data.frame': 7 obs. of 8 variables:
#> ..$ nam: chr [1:7] "vc" "mvi" "mnr" "mci" ...
#> ..$ vc : chr [1:7] "vc" "mvi" "vc" "mci" ...
#> ..$ mvi: chr [1:7] "mvi" "mvi" "err" "mci" ...
#> ..$ mnr: chr [1:7] "vc" "err" "mnr" "mci" ...
#> ..$ mci: chr [1:7] "mci" "mci" "mci" "mci" ...
#> ..$ mbd: chr [1:7] "err" "err" "err" "err" ...
#> ..$ mir: chr [1:7] "vc" "err" "mir" "mci" ...
#> ..$ mbi: chr [1:7] "vc" "err" "mnr" "mci" ...
#> $ booklets :'data.frame': 3 obs. of 4 variables:
#> ..$ booklet: chr [1:3] "booklet1" "booklet2" "booklet3"
#> ..$ block1 : chr [1:3] "bl1" "bl4" "bl3"
#> ..$ block2 : chr [1:3] "bl2" "bl3" "bl1"
#> ..$ block3 : chr [1:3] "bl3" "bl2" "bl4"
#> $ blocks :'data.frame': 30 obs. of 3 variables:
#> ..$ subunit : chr [1:30] "I01" "I02" "I03" "I04" ...
#> ..$ block : chr [1:30] "bl1" "bl1" "bl1" "bl1" ...
#> ..$ subunitBlockPosition: chr [1:30] "1" "2" "3" "4" ...SPSS Data
To get the input from SPSS data files, readSpss() reads
the data into R and converts all variables to the class
character. The function needs the name of the SPSS data
file and has the option to specify some more attributes. For more
information use the help function ?readSpss.
#readSpss(system.file("extdata", "booklet1.sav", package = "eatPrep"))
#readSpss(system.file("extdata", "booklet2.sav", package = "eatPrep"))
#readSpss(system.file("extdata", "booklet3.sav", package = "eatPrep"))Merkmalsauszug
The software “IQB Item-DB” produces Excel files named “Merkmalsauszug” using information stored in the IQB Databases. The file is expected to have the sheets: “Itemmerkmale”, “Aufgabenmerkmale”. The order doesn’t matter. The file contains information about the attributes of the items and exercises.
The function readMerkmalXlsx() reads the Excel file into
R and will produce a warning if any sheets are missing or wrongly
specified. It needs one character string (filename) containing path,
name and extension of the Excel file (.xlsx) produced by IQB Item-DB. By
default, the Item-ID is created without converting numbers to lowercase
letters (tolcl) and a merged data frame containing both
“Itemmerkmale” and “Aufgabenmerkmale” will be created
(alleM).
The function returns a list of data frames containing Itemmerkmale, Aufgabenmerkmale and AlleMerkmale (optional).
filename <- system.file("extdata", "itemmerkmale.xlsx", package = "eatPrep")
readMerkmalXlsx(filename, tolcl = FALSE, alleM = TRUE)
#> Reading sheet 'Aufgabenmerkmale'.
#> Reading sheet 'Itemmerkmale'.
#> Data frame 'AlleMerkmale' has been created.
#> $Aufgabenmerkmale
#> Aufgabe Zeit.A AufgID AufgTitel
#> 1 Animals: Weight of a duck 0:00 Animals: Weight of a duck NA
#> 2 Animals: Weight of a horse 0:00 Animals: Weight of a horse NA
#>
#> $Itemmerkmale
#> Aufgabe Teilaufgabe Item Zeit.I Anforderungsbereich.MaP
#> 1 Animals: Weight of a duck A 01 0:00 I
#> 2 Animals: Weight of a horse A 01 0:00 I
#> 3 Animals: Weight of a horse B 02 0:00 II
#> Bildungsstandards.MaP.allgemeine Itemtyp.5er
#> 1 1a GA - Geschlossen Ankreuzen
#> 2 <NA> GA - Geschlossen Ankreuzen
#> 3 <NA> GA - Geschlossen Ankreuzen
#> AufgID AufgTitel ItemID
#> 1 Animals: Weight of a duck NA Animals: Weight of a duck01
#> 2 Animals: Weight of a horse NA Animals: Weight of a horse01
#> 3 Animals: Weight of a horse NA Animals: Weight of a horse02
#>
#> $AlleMerkmale
#> AufgID Aufgabe Teilaufgabe Item
#> 1 Animals: Weight of a duck Animals: Weight of a duck A 01
#> 2 Animals: Weight of a horse Animals: Weight of a horse A 01
#> 3 Animals: Weight of a horse Animals: Weight of a horse B 02
#> ItemID Zeit.I Anforderungsbereich.MaP
#> 1 Animals: Weight of a duck01 0:00 I
#> 2 Animals: Weight of a horse01 0:00 I
#> 3 Animals: Weight of a horse02 0:00 II
#> Bildungsstandards.MaP.allgemeine Itemtyp.5er Zeit.A
#> 1 1a GA - Geschlossen Ankreuzen 0:00
#> 2 <NA> GA - Geschlossen Ankreuzen 0:00
#> 3 <NA> GA - Geschlossen Ankreuzen 0:00Checks
After reading in all the data, we need to check, whether it is in the right format and all codes and missing codes were assigned correctly.
checkInputList
checkInputList() checks whether the object
inputList has the required form. It has two arguments: the
inputList and mistypes. The
mistypes argument defines the missing types that should be
checked in valueRecode and valueType.
data(inputList)
checkInputList(inputList)The xlsx-file produced by ‘ZKDaemon’ is expected to have the
following sheets: “units”, “subunits”, “values”, “unitrecoding”,
“sav-files”, “params”, “aggregate-missings”, “itemproperties”,
“propertylabels”, “booklets”, and “blocks”. readDaemonXlsx
will produce a warning if any sheets are missing or wrongly specified.
checkInputList performs many additional checks to this.
checkData
checkData() checks data frames for missing or duplicated
entries in the ID variable, persons and/or variables without valid
codes, and generally invalid codes. The results of that check will be
written in the console.
The function needs a data frame to be checked (dat) and
its name (datnam), especially if there are multiple data
frames (e.g. a list of data frames). Furthermore it needs data frames
with code, subunit and unit information (values,
subunits and units), and a string for an ID
column name (ID). You can turn off printing information to the console
with verbose.
checkData(dat, datnam, values, subunits, units, ID = NULL, verbose = TRUE)Examples of data frames for values,
subunits and units can be found by typing
inputList in the console.
checkDesign
checkDesign() checks whether a data frame corresponds to
a particular rotated block design, i.e. whether all persons have valid
codes on all items they were presented with and one consistent missing
code for all items they were not presented with.
The function also needs a data frame to be checked
(dat), as well as data frames containing information on the
number and column names of blocks in each booklet
(booklets), on the names of subunits and their order within
each block (blocks) and about which participant worked on
which booklet (rotation). sysMis specifies the
missing code for items that were not administered to a participant and
id indicates the name of the participant identifier
variable in dat. This is needed, when you perform
checkDesign() on a data frame that is the result of
recodeData() which renames subunits according
to subunitRecoded in subunits sheet.
subunits is an optional argument to identify the names
of recoded subunits. And you can turn off printing information with
verbose.
checkDesign(dat, booklets, blocks, rotation, sysMis="NA", id="ID", subunits = NULL, verbose = TRUE)inputDat and inputList are examples on how
these data frames are supposed to look like. When you copy and paste the
following code in your console, you can look at the data frames.
Merging, Recoding, Aggregating, Scoring
Merging Data
Now that we checked that the data frames meet our requirements, we
can merge a list of data frames into a single data frame by using
mergeData(). For that we need a list of data frames, like
inputDat which contains data of three booklets. The
function returns a data frame containing unique cases and unique
variables. All cases and all variables from the original data sets will
be kept and matched.
mergeData() provides detailed diagnostics about value
mismatches. If two identically named columns in two data sets do not
have identical values, NAs are replaced by valid codes stemming from the
other data set(s) and if two different valid values are found, the first
value will be kept and the other dropped, and the user will be informed
about the mismatch. Additionally, NA resulting from the
merge (e.g., in repeated block designs) can be replaced with a custom
character missing to facilitate future data preparation of the merged
data set. See Table Missing Types for
standard missing types for other functions in the eatPrep
package.
When merging data frames with this function you need to specify at
least two arguments: newID and datList.
newID has to be a character vector length one indicating
the name of the identifier variable (ID) in the merged data set and/or
the name of the ID in every data frame in datList, if not
specified differently in oldIDs. datList is
the list of data frames to be merged, e.g. inputDat.
mergedDataset <- mergeData(newID = "ID", datList = inputDat)
#> Start merging of dataset 1.
#> Start merging of dataset 2.
#> Start merging of dataset 3.
str(mergedDataset)
#> 'data.frame': 300 obs. of 32 variables:
#> $ ID : chr "person100" "person101" "person102" "person103" ...
#> $ hisei: num 49 NA 57 32 59 56 55 47 NA 50 ...
#> $ I01 : chr "1" "9" "3" "2" ...
#> $ I02 : chr "4" "4" "4" "4" ...
#> $ I03 : chr "1" "2" "2" "3" ...
#> $ I04 : chr "2" "2" "3" "1" ...
#> $ I05 : chr "0" "0" "0" "1" ...
#> $ I06 : chr "0" "9" "0" "1" ...
#> $ I07 : chr "2" "2" "2" "3" ...
#> $ I08 : chr "0" "1" "0" "0" ...
#> $ I09 : chr "2" "2" "2" "3" ...
#> $ I10 : chr "2" "1" "1" "4" ...
#> $ I11 : chr "4" "1" "3" "2" ...
#> $ I12a : chr "1" "0" "1" "0" ...
#> $ I12b : chr "0" "9" "0" "1" ...
#> $ I12c : chr "4" "1" "2" "4" ...
#> $ I13 : chr "9" "4" "9" "4" ...
#> $ I14 : chr "9" "4" "9" "1" ...
#> $ I15 : chr "9" "3" "9" "1" ...
#> $ I16 : chr "9" "2" "9" "2" ...
#> $ I17 : chr "9" "3" "9" "4" ...
#> $ I18 : chr "9" "1" "9" "1" ...
#> $ I19 : chr "9" "4" "9" "2" ...
#> $ I20 : chr "9" "1" "9" "1" ...
#> $ I21 : chr "9" "2" "9" "3" ...
#> $ I22 : chr NA NA NA NA ...
#> $ I23 : chr NA NA NA NA ...
#> $ I24 : chr NA NA NA NA ...
#> $ I25 : chr NA NA NA NA ...
#> $ I26 : chr NA NA NA NA ...
#> $ I27 : chr NA NA NA NA ...
#> $ I28 : chr NA NA NA NA ...Furthermore, you can specify some more arguments, but they have
default options, so you don’t need to. Here is an example where the IDs
are changed via oldIDs and where NAs are replaced by “mbd”
(missing by design). For more information use the help function
?mergeData.
mergedDataset2 <- mergeData(newID = "idstud", datList = inputDat, oldIDs = c("ID", "ID", "ID"), addMbd = TRUE)
#> Start merging of dataset 1.
#> Start merging of dataset 2.
#> Start merging of dataset 3.
#> Start adding mbd according to data pattern.
str(mergedDataset2)
#> 'data.frame': 300 obs. of 32 variables:
#> $ idstud: chr "person100" "person101" "person102" "person103" ...
#> $ hisei : num 49 NA 57 32 59 56 55 47 NA 50 ...
#> $ I01 : chr "1" "9" "3" "2" ...
#> $ I02 : chr "4" "4" "4" "4" ...
#> $ I03 : chr "1" "2" "2" "3" ...
#> $ I04 : chr "2" "2" "3" "1" ...
#> $ I05 : chr "0" "0" "0" "1" ...
#> $ I06 : chr "0" "9" "0" "1" ...
#> $ I07 : chr "2" "2" "2" "3" ...
#> $ I08 : chr "0" "1" "0" "0" ...
#> $ I09 : chr "2" "2" "2" "3" ...
#> $ I10 : chr "2" "1" "1" "4" ...
#> $ I11 : chr "4" "1" "3" "2" ...
#> $ I12a : chr "1" "0" "1" "0" ...
#> $ I12b : chr "0" "9" "0" "1" ...
#> $ I12c : chr "4" "1" "2" "4" ...
#> $ I13 : chr "9" "4" "9" "4" ...
#> $ I14 : chr "9" "4" "9" "1" ...
#> $ I15 : chr "9" "3" "9" "1" ...
#> $ I16 : chr "9" "2" "9" "2" ...
#> $ I17 : chr "9" "3" "9" "4" ...
#> $ I18 : chr "9" "1" "9" "1" ...
#> $ I19 : chr "9" "4" "9" "2" ...
#> $ I20 : chr "9" "1" "9" "1" ...
#> $ I21 : chr "9" "2" "9" "3" ...
#> $ I22 : chr "mbd" "mbd" "mbd" "mbd" ...
#> $ I23 : chr "mbd" "mbd" "mbd" "mbd" ...
#> $ I24 : chr "mbd" "mbd" "mbd" "mbd" ...
#> $ I25 : chr "mbd" "mbd" "mbd" "mbd" ...
#> $ I26 : chr "mbd" "mbd" "mbd" "mbd" ...
#> $ I27 : chr "mbd" "mbd" "mbd" "mbd" ...
#> $ I28 : chr "mbd" "mbd" "mbd" "mbd" ...Recoding Data
After importing the data and making sure it has the right format, the next step is to adjust the missing values.
First, you recode data sets with several kinds of missing values. For
that, you need recode information with special consideration of missing
values. See collapseMissings() for supported types of
missing values. recodeData() recodes the specified data
frames and will give warnings if missing or incomplete recode
information is found. Values without recode information will not be
recoded.
Examples of data frames values and subunits
can be found when copy-pasting the following code into your console:
inputList$values
inputList$subunitsrecodeData() uses the recode information from those two
data frames and recodes the variables on dat accordingly.
The columns will be named according to the specifications in
subunits$subunitRecoded, if subunits is not
provided, item names will not be changed for recoded items.
datRec <- recodeData(dat = inputDat[[1]], values = inputList$values, subunits = inputList$subunits, verbose = TRUE)
#>
#> Found no recode information for variable(s):
#> ID, hisei.
#> This/These variable(s) will not be recoded.
#>
#> Variables... I01, I02, I03, I04, I05, I06, I07, I08, I09, I10, I11, I12a, I12b, I12c, I13, I14, I15, I16, I17, I18, I19, I20, I21
#> ...have been recoded.
str(datRec)
#> 'data.frame': 100 obs. of 25 variables:
#> $ ID : chr "person100" "person101" "person102" "person103" ...
#> $ hisei: chr "49" NA "57" "32" ...
#> $ I01R : chr "0" "mbi" "1" "0" ...
#> $ I02R : chr "0" "0" "0" "0" ...
#> $ I03R : chr "0" "1" "1" "0" ...
#> $ I04R : chr "1" "1" "0" "0" ...
#> $ I05R : chr "0" "0" "0" "1" ...
#> $ I06R : chr "0" "mbi" "0" "1" ...
#> $ I07R : chr "0" "0" "0" "0" ...
#> $ I08R : chr "0" "1" "0" "0" ...
#> $ I09R : chr "0" "0" "0" "0" ...
#> $ I10R : chr "1" "0" "0" "0" ...
#> $ I11R : chr "0" "0" "1" "0" ...
#> $ I12aR: chr "1" "0" "1" "0" ...
#> $ I12bR: chr "0" "mbi" "0" "1" ...
#> $ I12cR: chr "1" "0" "0" "1" ...
#> $ I13R : chr "mbi" "0" "mbi" "0" ...
#> $ I14R : chr "mbi" "0" "mbi" "0" ...
#> $ I15R : chr "mbi" "0" "mbi" "1" ...
#> $ I16R : chr "mbi" "0" "mbi" "0" ...
#> $ I17R : chr "mbi" "0" "mbi" "0" ...
#> $ I18R : chr "mbi" "1" "mbi" "1" ...
#> $ I19R : chr "mbi" "1" "mbi" "0" ...
#> $ I20R : chr "mbi" "0" "mbi" "0" ...
#> $ I21R : chr "mbi" "0" "mbi" "1" ...Aggregating Data
After recoding missing values, the subunits are combined one by one
via aggregateData().
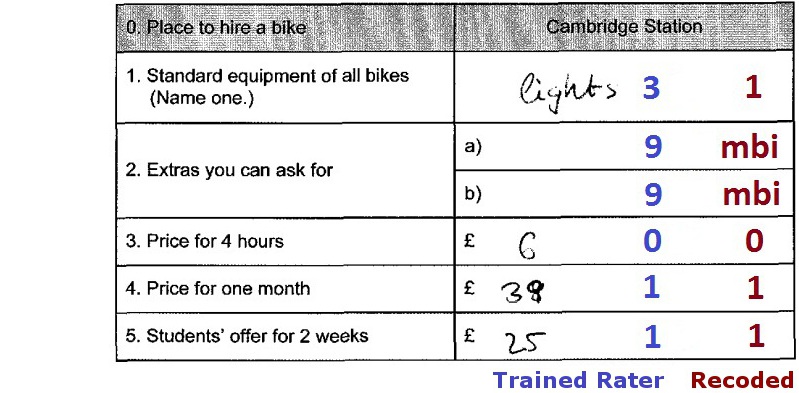
This function needs three data frames: one containing the data to be
aggregated (dat), one containing the subunit information
(subunits), and one containing the unit information
(units).
| vc | valid code |
| colnames | missing types to be aggregated |
| rownames | missing types to be aggregated |
Optionally, you can specify how missing values should be aggregated
(aggregatemissings) like in the example, or which column
names the returned data frame should have (recodedData),
for instance. Type ?aggregateData into your console to
learn more.
am <- matrix(c(
"vc" , "mvi", "vc" , "mci", "err", "vc" , "mbi", "err",
"mvi", "mvi", "err", "mci", "err", "err", "err", "err",
"vc" , "err", "mnr", "mci", "err", "mir", "mnr", "err",
"mci", "mci", "mci", "mci", "err", "mci", "mci", "err",
"err", "err", "err", "err", "mbd", "err", "err", "err",
"vc" , "err", "mir", "mci", "err", "mir", "mir", "err",
"mbi", "err", "mnr", "mci", "err", "mir", "mbi", "err",
"err", "err", "err", "err", "err", "err", "err", "err" ),
nrow = 8, ncol = 8, byrow = TRUE)
dimnames(am) <-
list(c("vc" ,"mvi", "mnr", "mci", "mbd", "mir", "mbi", "err"),
c("vc" ,"mvi", "mnr", "mci", "mbd", "mir", "mbi", "err"))
# using datRec from the chapter "recodeData()"
datAggr <- aggregateData(datRec, inputList$subunits, inputList$units,
aggregatemissings = am, rename = TRUE, recodedData = TRUE,
suppressErr = TRUE, recodeErr = "mci", verbose = TRUE)
#> All aggregation rules will be defaulted to 'SUM', because no other type is currently supported.
#> Found 20 unit(s) with only one subunit in 'dat'. This/these subunit(s) will not be aggregated and renamed to their respective unit name(s).
#> 1 units were aggregated: I12.Scoring Data
The next step is to score a data set with special consideration of
missing values. The function scoreData() is very similar to
recodeData(), but with a few defaults that are better
suited for scoring. scoreData() will give warnings when
incomplete scoring information is found. Values without scoring
information will not be scored.
Again, you need to specify three data frames. One data frame that you
get after completing the steps you did so far or by using
automateDataPreparation (dat), one with
information about the scoring of units (unitrecodings), and
one with subunit information (subunits). Examples for the
last two data frames can be found via inputList.
prepDat <- automateDataPreparation (inputList = inputList, datList = inputDat,
readSpss = FALSE, checkData=FALSE, mergeData = TRUE, recodeData=TRUE,
aggregateData=TRUE, scoreData= FALSE, writeSpss=FALSE, verbose = TRUE)
datSco <- scoreData(prepDat, inputList$unitRecodings, inputList$subunits,
verbose = TRUE)
#> ✔ 1 unit was scored: `I12`.mnrCoding
Then you can convert missing responses coded as “missing by
intention” (mbi) at the end of a block of items to “missing
not reached” (mnr) via mnrCoding(). The
function returns a data frame with “missing not reached” coded as
mnr. For each person with at least one mnr in
the returned data set the names of recoded variables are given as an
attribute to dat.
For that to work you need to specify the following arguments, as they don’t have any default settings:
| Argument | Explanation |
|---|---|
| dat | A data set. Missing by intention needs to be coded
mbi
|
| pid | Name or column number of the identifier (ID) variable in
dat
|
| rotation.id | A character vector of length 1 indicating the column name of the
test booklet identifier in dat
|
| blocks | A data frame containing the sequence of subunits in each block in
long format. The column names need to be subunit,
block, subunitBlockPosition
|
| booklets | A data frame containing the sequence of blocks in each booklet in
wide format. The column names need to be booklet,
block1, block2, block3, … |
| breaks | Number of blocks after which mbi shall be recoded to
mnr, e.g., c(1,2) to specify breaks after the
first and second block |
There are more arguments with default values which you can specify, but don’t have to.
nMbi, for instance, specifies the number of subunits at
the end of a block that need to be coded mbi. The default
is 2, i.e. if the last and second to last subitem in a block are coded
mbi, both subunits, as well as the preceding subunits coded
mbi, will be recoded to mnr. If
nMbi is larger than the number of subunits in a given
block, no subitem in this block will be recoded. If all subunits in a
block are coded mbi, none of them will be recoded to
mnr. nMbi needs to be > 0.
subunits has the default NULL, but when you
specify a data frame, mnrCoding() expects to find the
recoded subunits in dat.
Examples for the data frames booklets,
blocks, rotation and subunits can
be found via inputList. Here is an example use case of
mnrCoding(). The first two functions
(automateDataPreparation() and mergeData())
create the data frame dat for mnrCoding().
prepDat <- automateDataPreparation(inputList = inputList,
datList = inputDat, readSpss = FALSE, checkData=FALSE,
mergeData = TRUE, recodeData=TRUE, aggregateData=FALSE,
scoreData= FALSE, writeSpss=FALSE, verbose = TRUE)
prepDat2 <- mergeData("ID", list(prepDat, inputList$rotation))
mnrDat <- mnrCoding(dat = prepDat2, pid = "ID",
booklets = inputList$booklets, blocks = inputList$blocks,
rotation.id = "booklet", breaks = c(1, 2),
subunits = inputList$subunits, nMbi = 2, mbiCode = "mbi",
mnrCode = "mnr", invalidCodes = c("mbd", "mir", "mci"),
verbose = TRUE)
#> ...identifying items in data (reference is blocks$subunit)
#> Variables in data not recognized as items:
#> ID, hisei, booklet
#> If some of these excluded variables should have been identified as items (and thus be used for mnr coding) check 'blocks', 'subunits', 'dat'.
#> ...identifying items with no mbi-codes ('mbi'):
#> I04R, I08R
#> If you expect mbi-codes on these variables check your data and option 'mbiCode'
#> mnr statistics:
#> mnr cells: 553
#> unique cases with at least one mnr code: 89
#> unique items with at least one mnr code: 16
#> unique cases ('ID') per booklet and booklet section (0s omitted):
#> booklet booklet.section N.ID
#> 1 booklet1 2 11
#> 2 booklet1 3 28
#> 3 booklet2 1 28
#> 4 booklet2 2 11
#> 5 booklet2 3 1
#> 6 booklet3 3 31
#> start recoding (item-wise)
#> done
#> elapsed time: 0.1 secsType ?mnrCoding into your console to learn more.
Wrapper
Instead of using all the individual functions as described above, you
can also use the wrapper function automateDataPreparation,
which contains most of the described functions. Here is an overview over
the arguments that call the corresponding function.
| Argument | Corresponding Function |
|---|---|
readSpss |
readSpss() |
checkData |
checkData() |
mergeData |
mergeData() |
recodeData |
recodeData() |
recodeMnr |
mnrCoding() |
aggregateData |
aggregateData() |
scoreData |
scoreData() |
collapseMissings |
collapseMissings() |
writeSpss |
writeSpss() |
But there are many more arguments that may need to be considered. For
more information see ?automateDataPreparation.
Here is an example use case and its output.
automateDataPreparation returns a data frame resulting from
the final data preparation step.
data(inputList)
data(inputDat)
preparedData <- automateDataPreparation(inputList = inputList,
datList = inputDat, path = getwd(),
readSpss = FALSE, checkData = TRUE, mergeData = TRUE,
recodeData = TRUE, recodeMnr = TRUE, breaks = c(1,2),
aggregateData = TRUE, scoreData = TRUE,
writeSpss = FALSE, verbose = TRUE)
#> Starting automateDataPreparation 2026-05-13 18:03:33.85443
#>
#> Check data...
#>
#> Checking dataset booklet1
#> Only valid codes in ID variable.
#> No duplicated entries in ID variable.
#> No duplicated variable names.
#> Found no variable information about variable(s) hisei. This/These variables will not be checked for missings and invalid codes.
#> Found no invalid codes.
#>
#> Checking dataset booklet2
#> Only valid codes in ID variable.
#> No duplicated entries in ID variable.
#> No duplicated variable names.
#> Found no variable information about variable(s) hisei. This/These variables will not be checked for missings and invalid codes.
#> Found no invalid codes.
#>
#> Checking dataset booklet3
#> Only valid codes in ID variable.
#> No duplicated entries in ID variable.
#> No duplicated variable names.
#> Found no variable information about variable(s) hisei. This/These variables will not be checked for missings and invalid codes.
#> Found no invalid codes.
#>
#> Start merging.
#> Start merging of dataset 1.
#> Start merging of dataset 2.
#> Start merging of dataset 3.
#> Start adding mbd according to data pattern.
#>
#> Start recoding.
#>
#> Found no recode information for variable(s):
#> ID, hisei.
#> This/These variable(s) will not be recoded.
#>
#> Variables... I01, I02, I03, I04, I05, I06, I07, I08, I09, I10, I11, I12a, I12b, I12c, I13, I14, I15, I16, I17, I18, I19, I20, I21, I22, I23, I24, I25, I26, I27, I28
#> ...have been recoded.
#>
#> Start recoding Mbi to Mnr.
#> ...identifying items in data (reference is blocks$subunit)
#> Variables in data not recognized as items:
#> ID, booklet, hisei
#> If some of these excluded variables should have been identified as items (and thus be used for mnr coding) check 'blocks', 'subunits', 'dat'.
#> ...identifying items with no mbi-codes ('mbi'):
#> I04R, I08R
#> If you expect mbi-codes on these variables check your data and option 'mbiCode'
#> mnr statistics:
#> mnr cells: 553
#> unique cases with at least one mnr code: 89
#> unique items with at least one mnr code: 16
#> unique cases ('ID') per booklet and booklet section (0s omitted):
#> booklet booklet.section N.ID
#> 1 booklet1 2 11
#> 2 booklet1 3 28
#> 3 booklet2 1 28
#> 4 booklet2 2 11
#> 5 booklet2 3 1
#> 6 booklet3 3 31
#> start recoding (item-wise)
#> done
#> elapsed time: 0.1 secs
#>
#> Start aggregating
#> Since inputList$aggrMiss exists, this will be used instead of default.
#> All aggregation rules will be defaulted to 'SUM', because no other type is currently supported.
#> Found 27 unit(s) with only one subunit in 'dat'. This/these subunit(s) will not be aggregated and renamed to their respective unit name(s).
#> 1 units were aggregated: I12.
#>
#> Start scoring.
#> ✔ 1 unit was scored: `I12`.
#>
#> No SPSS-File has been written.
#>
#> Missings are UNcollapsed.
#> automateDataPreparation terminated successfully! 2026-05-13 18:03:34.102173Additional Diagnostics and Rater Tools
The functions above cover the main data preparation pipeline.
eatPrep also contains tools for category diagnostics, rater
agreement, IDM cut scores and semi-manual recoding of flagged
subsets.
Category Discrimination
catPbc() calculates category-level point-biserial
correlations. It compares a merged raw data set with the corresponding
recoded data set and returns frequencies and correlations for each item
category. This is useful for finding recoding mistakes or suspicious
item categories before scaling.
datRaw <- mergeData(newID = "ID", datList = inputDat, addMbd = TRUE)
#> Start merging of dataset 1.
#> Start merging of dataset 2.
#> Start merging of dataset 3.
#> Start adding mbd according to data pattern.
datRecPbc <- recodeData(datRaw, values = inputList$values,
subunits = inputList$subunits, verbose = FALSE)
pbcs <- catPbc(datRaw, datRecPbc, idRaw = "ID", idRec = "ID",
context.vars = "hisei", values = inputList$values,
subunits = inputList$subunits)
head(pbcs)
#> item cat n freq freq.rel catPbc recodevalue subunitType
#> 1 I01 1 200 27 0.135 -0.05898175 0 1
#> 2 I01 2 200 84 0.420 -0.35201750 0 1
#> 3 I01 3 200 35 0.175 0.15887365 1 1
#> 4 I01 8 200 48 0.240 0.26930074 mir 1
#> 5 I01 9 200 6 0.030 0.10854342 mbi 1
#> 6 I02 1 200 19 0.095 0.13417790 1 1evalPbc() evaluates the output of catPbc()
against configurable thresholds. It returns an empty list if no
problematic categories are found; otherwise it returns item names
grouped by the type of potential problem.
evalPbc(pbcs)
#> ✖ The attractors (score 1) of the following 1 item were chosen with a frequency
#> of zero: I14. This should not happen. Please check.
#> ! The distractors (score 0) of the following 1 item were chosen with a
#> frequency of zero: I14_7. This may happen, but is probably not intended.
#> ✖ catPbcs for attractors (score 1) of the following 3 items are worrisome low (< 0.05) or missing: I12c:_0.01, I14:_NA, and I24:_-0.12
#> ✖ catPbcs for distractors (score 0) of the following 9 items are unexpectedly high (> 0.005): I02_4_0.11, I07_3_0.26, I07_4_0.03, I12c_3_0.03, I13_4_0.01, I14_4_0.06, I14_5_0.04, I15_3_0.03, I17_1_0.06, I17_3_0.04, I22_3_0.13, and I24_1_0.21
#> ! catPbcs for mistype 'mir' of the following 3 items are relatively high (>
#> 0.07): I01_8_0.27, I03_8_0.29, and I16_8_0.08
#> ! catPbcs for mistype 'mbi' of the following 8 items are relatively high (>
#> 0.07): I01_9_0.11, I02_9_0.15, I16_9_0.14, I17_9_0.16, I18_9_0.19, I20_9_0.14,
#> I21_9_0.15, and I23_9_0.17
#> ℹ For a list of problematic items, save the `output` of this function and
#> return the item names as a vector:
#> • `output$zeroFreqAtt`
#> • `output$zeroFreqDis`
#> • `output$lowMisPbcAtt`
#> • `output$highPbcDis`
#> • `output$highPbcMis$mir`
#> • `output$highPbcMis$mbi`Rater Agreement
meanAgree() and meanKappa() summarize
pairwise agreement between several raters. The input is a wide data
frame with persons in rows and raters in columns. If your rater data are
in long format, you can reshape them first.
Pseudo Raters
make.pseudo() reduces ratings from multiple real raters
to a smaller number of pseudo raters. This is useful when later
processing steps require a fixed number of ratings per response.
Optional arguments such as alwaysPrefer,
alwaysNeglect and item.groups let you control
which raters should be preferred and whether linked variables should
stay with the same pseudo rater where possible.
oneRater <- make.pseudo(datLong = rater, idCol = "id", varCol = "variable",
codCol = "rater", valueCol = "value", n.pseudo = 1,
verbose = TRUE)
#> N.persons: 1287
#> N.vars: 7
#> N.coder: 4
#> coders per response: minimum 2, maximum 2
#> responses with multiple ratings: 2100 of 2100 (100 %)
head(oneRater)
#> id rater variable value index
#> P0002 P0002 John V01 1 P0002_V01
#> P0009 P0009 Carol V01 0 P0009_V01
#> P0010 P0010 Edward V01 0 P0010_V01
#> P0020 P0020 John V01 9 P0020_V01
#> P0022 P0022 Carol V01 1 P0022_V01
#> P0024 P0024 Dolores V01 0 P0024_V01IDM Cut Scores
computeCutsIDM() calculates cut scores for Item
Descriptor Matching (IDM). The data set must contain item difficulty
estimates and one or more numeric rater columns. The
boundaries argument determines the requested cut scores,
for example four boundaries for five performance levels.
idmDat <- data.frame(
est = seq(-2, 2, length.out = 8),
Rater1 = c(1, 1, 2, 2, 3, 4, 4, 5),
Rater2 = c(1, 2, 2, 3, 3, 4, 5, 5)
)
cuts <- computeCutsIDM(idmDat)
cuts$cuts_summary
#> # A tibble: 1 × 4
#> cut12 cut23 cut34 cut45
#> <dbl> <dbl> <dbl> <dbl>
#> 1 -1.14 -1.11e-16 0.857 1.71plotCutsIDM() visualizes the raw ratings, moving
averages, monotonized curves and resulting cut scores.
plotCutsIDM(cuts)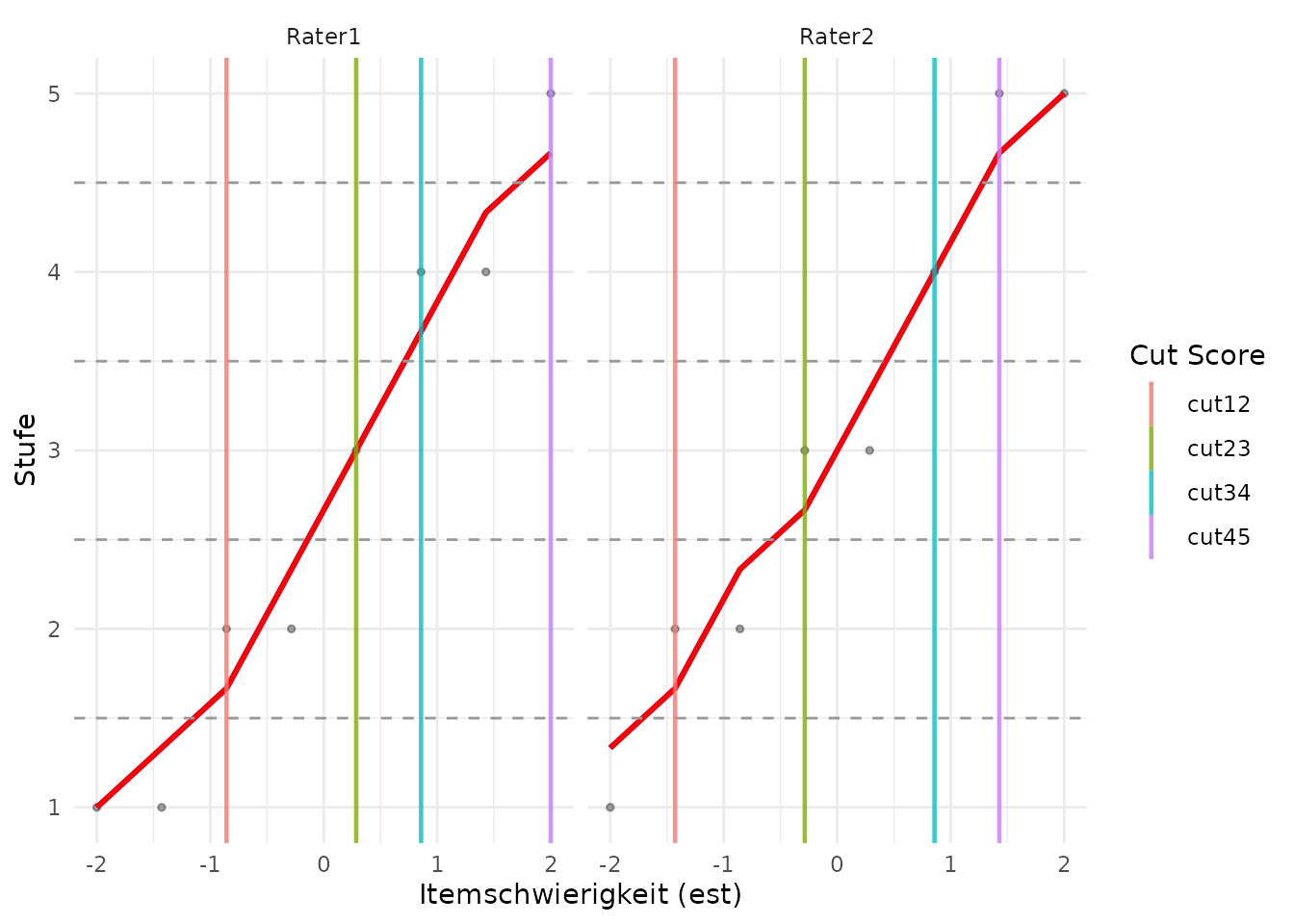
Visual Subset Recoding
visualSubsetRecode() supports interactive inspection and
recoding of flagged subsets. For example, if a group of students was
affected by a technical issue, you can display the relevant responses
and decide whether to recode them to a value such as mci.
Because the function opens an interactive review flow, it is not
evaluated in this vignette.
subsetInfo <- data.frame(
ID = c("person100", "person101", "person102"),
datCols = c("I01", "I02", "I03")
)
datVisRec <- visualSubsetRecode(dat = preparedData, subsetInfo = subsetInfo,
ID = "ID", toRecodeVal = "mci")Export
After the data preparation is done, you can prepare the data sets for
exporting to then use them in the packages eatModel and
eatGADS.
Collapsing Data
The last step is to recode character missings to numerical types with
collapseMissings(), usually to 1, 0 or NA, but other values
are also possible. All variables need to be converted to
numeric, as well, so you can pass the data set to
eatModel afterwards.
You can use the data frame that is returned by
recodeData(), which we called datRec earlier
(dat). But its main purpose is to finish the scored data
set as it comes out of scoreData() or
automateDataPreparation().
missing.rule is a list containing information which
character missings should be converted to 0 or
NA. It has default settings, but you can adapt them if
needed. Type ?collapseMissings into your console to learn
more.
datColMis <- collapseMissings(datRec)After doing that you should have a data frame that you can now use
for further processing with the eatModel package.
Export SPSS
With writeSpss() you can create a .txt file and a SPSS
syntax file, which can then be used in SPSS. You need the four data
frames dat, values, subunits and
units, the last three are usually part of the
inputList. You also need to specify the file names and
where to save those new files.
The function creates and saves those files, but the actual return
values for this function is NULL.
Prepare for eatGADS
prep2GADS converts the data frame that we prepared to an
eatGADS object for further use in the package eatGADS. It
uses the meta data stored in inputList, so it needs the
inputList as an argument, as well as our data frame
dat, that we have been preparing.
Here you can see two examples for creating eatGADS objects. The first
example exports scored data (trafoType = “scored”), which
usually has 0/1 values and mistypes like mbi/mbo etc. The second example
exports the raw data (trafoType = “raw”), including the
original values and subunits.
data(inputDat)
data(inputList)
# for GADSobj1
prepDatScored <- automateDataPreparation(inputList = inputList, datList = inputDat,
readSpss = FALSE, checkData=FALSE, mergeData = TRUE, recodeData=TRUE,
aggregateData=TRUE, scoreData=TRUE, writeSpss=FALSE, verbose = TRUE)
# for GADSobj2
prepDatRaw <- automateDataPreparation(inputList = inputList, datList = inputDat,
readSpss = FALSE, checkData=FALSE, mergeData = TRUE,
recodeData=FALSE, aggregateData=FALSE, scoreData=FALSE,
writeSpss=FALSE, verbose = TRUE)
GADSobj1 <- prep2GADS(dat = prepDatScored, inputList = inputList[1:3], trafoType = "scored",
verbose=TRUE)
#>
#> ── Check: Variables without info
#> ℹ The following 1 variable is not in inputList ($units$unit) but in dataset,
#> its meta data will be set to NA: `hisei`
GADSobj2 <- prep2GADS(dat = prepDatRaw, inputList = inputList[1:3], trafoType = "raw",
verbose=TRUE)
#>
#> ── Check: Variables without info
#> ℹ The following 1 variable is not in inputList ($subunits$subunit or
#> $units$unit) but in dataset, its meta data will be set to NA: `hisei`